
Design and function of molecular machines
Lab closed in 2013.
What keeps cells and organisms alive are specific functions performed by highly organized macromolecular assemblies. Thomas Marlovits' research group wants to understand the architectural design of these ‘biological nanomachines’. The team explores such structures under normal and disease-related conditions. In order to elucidate the molecular mechanisms, the research group uses high-resolution three-dimensional electron microscopy to directly visualize molecular machines in action.
As a joint IMP-IMBA group leader Thomas Marlovits works for both IMBA and the partner institute IMP.
Research
Molecular Machines
Membrane-associated processes are a fundamental characteristic of all living cells because they ensure that the cells are able to effectively communicate with, and adapt to, their environment. Cells achieve this by either physically translocating molecules to the opposite site of a membrane, or by receiving, transmitting, and amplifying incoming signals. Our laboratory is interested in understanding the molecular mechanism underlying such processes. Specifically, we focus on machineries capable of translocating bacterial toxins into eukaryotic cells.


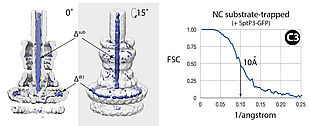

Microbial pathogenesis
Gram-negative pathogens such Yersinia, Shigella, Pseudomonas, enteropathogenic/enterohemorrhagic E. coli (EPEC/EHEC) and Salmonella in animals are causative agents for many diseases in animals and humans. The diseases have mild to deadly outcomes, and often originate as food-borne diseases. A crucial element of their pathogenicity are bacterial toxins (‘effectors’), which are delivered via the type III secretion system (a large membrane-embedded machinery) from the bacterium to its host cell. As a consequence, translocated effector proteins have the remarkable capacity to modulate various host-cell pathways, including endocytic trafficking, gene expression, programmed cell death, or cytoskeleton dynamics that induce membrane ruffling and subsequently render the host accessible to bacterial infection.
Unfolded protein transport across membranes?
Safe and directional transport of effector proteins across membranes is the hallmark function of all type III secretion. Our recent structural analysis (Schraidt & Marlovits, Science, 2010) of the injectisome, the most prominent cylindrical structure of the type III secretion system, revealed a potential secretion path through the central part of the membrane-embedded complex. However, the inner diameter of this path is too small to accommodate a fully folded effector protein, suggesting that either the injectisome must undergo large conformational changes during transport, or effector proteins must be unfolded.
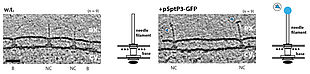
To investigate type III secretion of human pathogens, we focused on (1) determining the secretion path of injectisomes, (2) understanding the mechanism of transport, and (3) visualizing protein transport in situ. In order to address these questions, we first analyzed the requirements for substrate association with, transport through, and exit through the injectisome. Surprisingly, we found that the size and length of novel substrates does not exert a major impact on their secretability. We learned that the fusion of thermodynamically stable protein domains to otherwise secreted substrates does not influence successful engagement to the injectisome, but prevents complete transport across membranes. Such designed and trapped substrates are commonly associated with injectisomes.
We discovered that the substrates are inserted into the secretion path in polar fashion (N-terminal regions first), suggesting that other substrates with a similar domain organization follow the same principle. Our structural analysis of trapped substrates revealed that they are in an unfolded state during transport, suggesting that the type III-specific ATPase acts as an unfoldase. In contrast, injectisomes stay largely invariant during protein transport. To understand whether such behavior is in fact observed in situ, we performed cryo-electron tomography. This method permits the investigation of molecular structures within cells in a spatiotemporal manner and in a near-native state. For the first time we were able to visualize pathogenic type III secretion systems from Salmonella in action and - more generally - protein transport across several membranes.
By understanding the molecular mechanism of TTSS-mediated protein transport, we hope to provide a basis for the development of novel therapeutic strategies that will either inhibit its activity or modify the system for targeted drug delivery.
Team
- BRUNNER Matthias
- CHABLOZ Antoine
- CICCARELLI Luciano
- GÖSSWEINER-MOHR Nikolaus
- KOTOV Vadim
- MARLOVITS Thomas
- MAYR Julia
- MILETIC Sean
- VESPER Oliver
- WALD Jiri
- ZALESAK Jan
Selected Publications
(2015)
DiMaio, F., Song, Y., Li, X., Brunner, MJ., Xu, C., Conticello, V., Egelman, E., Marlovits, TC., Cheng, Y., Baker, D. (2015). Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat Methods. 12(4):361-5 (abstract)
(2014)
Radics, J., Königsmaier, L., Marlovits, TC. (2014). Structure of a pathogenic type 3 secretion system in action. Nat Struct Mol Biol. 21(1):82-7 (abstract)
(2011)
Schraidt, O., Marlovits, TC. (2011). Three-dimensional model of Salmonella's needle complex at subnanometer resolution. Science. 331(6021):1192-5 (abstract)
(2010)
Wagner, S., Königsmaier, L., Lara-Tejero, M., Lefebre, M., Marlovits, TC., Galán, JE. (2010). Organization and coordinated assembly of the type III secretion export apparatus. Proc Natl Acad Sci U S A. 107(41):17745-50 (abstract)
(2006)
Marlovits, TC., Kubori, T., Lara-Tejero, M., Thomas, D., Unger, VM., Galán, JE. (2006). Assembly of the inner rod determines needle length in the type III secretion injectisome. Nature. 441(7093):637-40 (abstract)