WELCOME TO THE BURGA LAB
Words. Words. I play with words, hoping that some combination, even a chance combination, will say what I want.
The sequencing of the human genome over fifteen years ago ignited my passion for genetics and began a scientific revolution that continues to expand. While most people will have access to their own genome in the near future, many fundamental questions remain unanswered. What makes individuals, populations and ultimately species different from each other? What are the genetic and molecular mechanisms responsible for the myriad of adaptations present in nature?How accurately can we predict our physical attributes, behaviors and health early in life?
Unfortunately, our growing ability to read genomes and catalog genetic variants does not match our ability to understand nature and make useful predictions for people. We believe this problem reflects our limited understanding of the architecture and evolution of complex traits.
Lab Mission
Our overall goal is to understand at the molecular level the forces driving the evolution of complex traits with a particular emphasis on the contribution of epistatic interactions (genetic modifiers) and genomic conflict (selfish elements). To do so, we combine our expertise in quantitative genetics, genomics, and developmental biology. We strive to find elegant molecular explanations to the big questions in genetics and evolutionary biology.
Projects
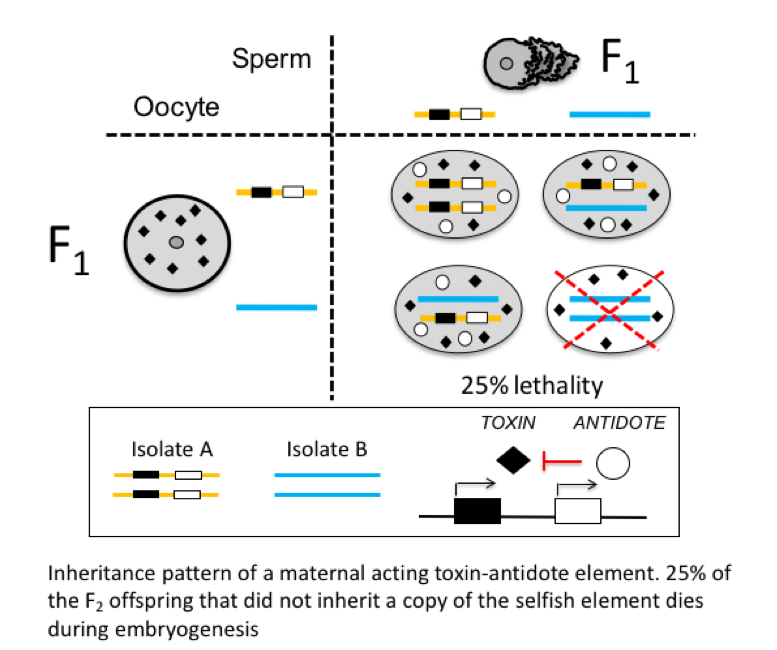
Toxin-Antidote Selfish Elements and Speciation
How could your parental genotype, including those alleles that you did not inherit, affect your biology? This process occurs in nematodes, where parental–zygotic genetic interactions can be the difference between life and death. The genes responsible for this unusual pattern of inheritance are selfish genetic elements propagating in wild populations exploiting their gene drive activity. These selfish elements subvert the laws of Mendelian segregation to promote their own inheritance. They are typically composed of two tightly linked genes: a toxin deposited in the gamete (oocyte or sperm) and its antidote expressed by the zygote.
After over 30 years of extensive research in C. elegans, one of the best-understood model organisms, we have identified the presence of two toxin-antidote selfish elements. I predict that more will be identified, which will stimulate new research areas. What are their molecular and functional mechanisms? How common are these elements in nature? In what species do they exist? How do they contribute to evolution and disease? These are some long-term questions I will address in my laboratory.
Genetic Modifiers in disease and evolution
One main factor that limits our ability to make accurate phenotypic predictions for individuals is the influence of the genetic background. Mutations do not occur in isolation, but in the context of thousands of other coding and non-coding variants that together constitute an individual’s genetic makeup. These natural variants segregate in populations and interact with disease-causing mutations to affect their outcome. For instance, they can increase or decrease the disease probability for an individual. Such variants are known as natural genetic modifiers. The presence of genetic modifiers has long been recognized in model organisms, but their effect has not received as much attention in humans. For example, very recent studies of cystic fibrosis and Huntington’s disease patients revealed the presence of large effect genetic modifiers of disease risk. This is remarkable, since both cystic fibrosis and Huntington’s disease are textbook examples of Mendelian inheritance. However, the vast majority of genetic modifiers in both humans and model organisms remain unknown.
How many natural genetic modifiers occur per gene? What is their molecular identity? What features do they share in common? Do these natural modifiers match those found by synthetic interactions screens?These are some of the questions we are addressing harnessing the power of CRISPR screens and standing natural genetic variation.
And More!
We are continuously exploring new areas of research, applying the latest genomics and molecular tools to study long standings questions in genetics and evolutionary biology.
Approach
Our lab combines experimental computational approaches and always welcomes candidates that want to work at their interface. Pipette by day, code by night!
Selected Publications
Ben-David, E., Pliota, P., Widen, SA (...) Burga, A., Kruglyak, L. (2021). Ubiquitous Selfish Toxin-Antidote Elements in Caenorhabditis Species. Curr Biol. 31(5):990-1001.e5
Burga, A., Ben-David, E., Lemus Vergara, T., Boocock, J., Kruglyak, L. (2019). Fast genetic mapping of complex traits in C. elegans using millions of individuals in bulk. Nat Commun. 10(1):2680